This post addresses the conundrum that Staph aureus is a different bug biochemically depending on its environment. The Merck Manual gives four sites of Staph aureus infections: the blood, , the skin, the lung, and the bone. What we know about Staph growing in standard culture media under aerobic conditions may not approximate metabolism in all locations in our bodies that it may infect. The functional groups that Staph aureus uses to sense oxygen can signal changes in gene transcription. These groups contain unpaired electrons and may also be antenna for pulsed electromagnetic frequencies. PEMF might be a non invasive way to muck up the signaling that Staph aureus uses to switch to anaerobic growth. And yes, extremely low frequency PEMF kills Staph aureus. We don’t know why. This post makes the case that O2 sensing might be part of it.
- In blood, and presumably the lung, 95-100% O2 saturation is considered healthy according to Medical News Today.
- Bone marrow is around 1-9% saturation according to the Johnson 2017 review of the literature. These authors consider bond to be hypoxic.
- basal epidermis is around 2.5% while hair follicles and sebaceous glands are even lower around 0.12% according to a literature review in Chettouh-Hammas and Gillon 2024. What’s more gene expression changes in going from traditional soy based broth to one chemically similar to skin. Costa 2024
This post is a continuation of killing faculatative anaerobes with very low frequency PEMF and a a sister post to Staph aureus in atopic dermatitis. What can the metabolic regulation of Staph aureus tell us about potential PEMF antenna? Just in going from a common culture broth to a skin like culture broth, some changes are seen in gene transcription of potential PEMF antenna proteins. The feature image suggests to expose culture tubes (or pickle jars) of bacteria to PEMF inside Helmholtz coils and then monitor their growth. Is there a a switch to O2 preference? The featured image is meant to suggest that O2 preference can be inferred by where bacteria grow in a container.
Extremely low frequency PEMF inhibits Staph aureus, table
One of the frustrating things about these studies is that they show promise but are not performed under conditions that resemble those in encountered in an actual patient. The authors use standard microbiological media, but do the ingredients change the proteins that the bug expresses? With that caveat, these are some nice examples of promising studies.
| medium | O2 | time | frequency | magnetic field | outcome | ref |
|---|---|---|---|---|---|---|
| Columbia 5% sheep blood broth | room | 37o C for 24h | 0-50 Hz | 0.5-10 mT | 5 mT, 20 Hz 29.2% ↓ CFU (P = 0.0376) | Obermeier 2009 |
| TY broth tryptone yeast | room | 25oC 24 min | 50Hz | 7.1 mT | 20% ↓ CFU p<0.05 | Fojt 2004 |
| Mueller-Hinton broth, protein rich | room | 37o C for 24 h | 20 Hz | 5 mT | PEMF + antibiotic 32.4% of gentamicin alone p=0.0002 | Matl 2011 |
| MHB, in Eppendorf tube | ? | 25oC 1-6hr | 20, 40, 50 | 2 & 4 mT | 6 hr, 4 mT, 20 Hz best. | Bayır 2015 |
The blood and lungs are two sites of Staph infections. What gets turned on or off in more anaerobic regions of our bodies like bone and lower levels of the skin? It is interesting to note that many commercial PEMF devices use 50 Hz. The Micro-Pulse M1 “beta 3 wave” program emits at 20 and 28 Hz. No… There is absolutely, positively no proof whatsoever in this world that this program is going to kill a nasty case of Staph aureus atopic dermatitis. I’m just saying there is potential to be investigated on top of standard of care.
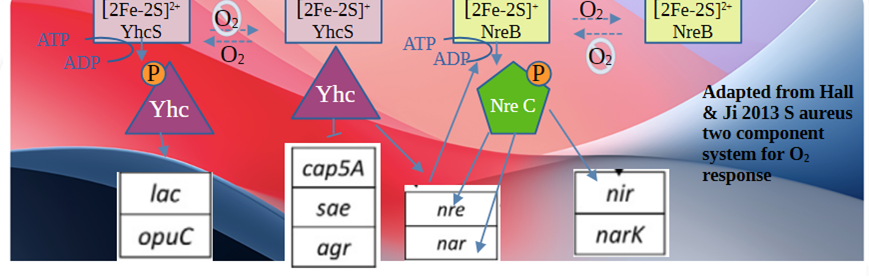
Hall and Ji 2013, what gets turned on and off with O2
This post will take on the same outline with comments on how this pertains to PEMF and changes in gene expression when going from a standard in vitro broth to a skin like growth.
Hall and Ji, Anaerobic metabolism of S. aureus
As a faculatative anaerobic bacterium, Staph aureus has the ability to reduce nitrate, instead of O2, as the terminal electron acceptor of the electron transport chain. Sugars and amino acids may be fermented and enter the TCA cycle to produce NADH for the electron transport chain. In the absence of O2, the nar and nir operons, which are responsible for nitrate and nitrite reduction, are up regulated. The authors didn’t comment on whether sulfate from sulfur containing amino acids oar nitrate from all amino aides was more important. Fumarate can also be a terminal electron acceptor as introduced in the 5G and microbiome post. It was asked what separated survivors from those killed by 5G PEMF.
Hall and Ji, Oxygen-Independent Regulation
Which of these sensors have the potential to be PEMF antenna? NAD+/NADH was mentioned a lot in the preamble to the key players.
- Rex—NAD/NADH sensor According to the Hall and Ji review, Rex transcription factors are kept in an active state when the radio of NAD+ to NADH is high, which happens when the electron transport chain and the export of H+ (proton motive force) out of the bacterium is high. Gene targets of Rex homodimer transcription factors include Idh1, ddh, adhE, pfl, ssrAB, and nar/nirR
- SsrAB SsrA is a histidine kinase that autophorphorylates in the presence of ATP. Its transcription factor target is SsrB, which promotes the transcription of ldh1, adhE, and the ATPase. SsrB inhibits the transcription of ndh, sucC, argD, citB and fumC.
- ArcR positively regulates the expression of the arcABDCR operon under anaerobic conditions, by binding to the consensus motif 5′-TGTGAN6TCACA-3′. Makhlin 2007 demonstrated that the ArcR Crp/Fnr family member controls the transcription of Arginine Deiminase Operon. They cited homology to a previous study that demonstrated redox sensitive formation of disulfide bonds in a Bacillus licheniformis close homolog (Wohlkönig 2004).
Even more so than people, bacteria behave differently in a nutrient rich versus poor environment. From a citizen scientist vantage, would PEMF change bacteria growth more, less or not at all if bean broth were spiked with beef broth, table sugar, a bit of blood?
Hall and Ji, Oxygen-Dependent Regulators
In the absence of an FNR protein, S. aureus has evolved two TCSs (two component systems) that are histidine kinases that sense environmental factors that are relayed to changes in gene transcription. They are responsible for sensing O2 and altering gene expression to use alternative terminal-electron acceptors or fermentation for energy production.
- NreABCNreB is the histidine kinase of the two component ssytem that has a Fe-S center. NreB/NreC activates the expression of the nitrate (narGHJI) and nitrite (nir) reductase operons, as well as the putative nitrate transporter gene narT (By similarity). NreC is the transcription. UniProt does not have much to say about NreA.
- AirSR AirS has a [2Fe-2S] cluster is essential for the histidine kinase activity. The oxidized cluster [2Fe-2S]2+ is in the fully active form of the kinase as opposed to the reduced [2Fe-2S]+ cluster. AirR is the transcription factor that controls transcription of the virulence factor sspABC operon and crtOPQMN operon needed for proeuction of staphyloxanthin.
- Both TCSs are wide spread throughout nonsporulating and sporulating Firmicutes suggesting a universal mechanism of O2 sensing for this phylum of eubacteria
So very low frequency PEMF can reduce the numbers of Staph aureus, what happens in low oxygen saturation would be really helpful information because so many sensors with unpaired electrons direct this bug to change its metabolism.
Costa 2024, Culture medium affects transcription in Staph aureus
The PEMF studies referenced in this post are promising but we really need to consider that the growth medium may affect the genes that are transcribed and thus the reaction to PEMF. Costa 2024 compared the gene expression in tryptic soy broth with a solution that was formulated to resemble human skin.
Costa 2024 summary of figures
- Fig 1, a link to the recipe for the Skin Like Medium. L-lactate, 20 mM and 1 mM glucose are the main sources of carbon. This mix contains 10 mM urea as a source of carbon and nitrogen. A B vitamin mix and amino acids were also used.
- Fig 2 presented growth, as measured by absorbance at 600nm in TSB and SLM. It might be nice to expose Staph aureus in liquid culture to PEMF at various culture densities.
- TSB was the reference and contains a digest of soy and casein. Compared to SLM, TSB contains peptides, a higher concentration of amino acids, phosphate, and glucose, and a lower concentration of sodium chloride.
- Fig 3 looks at differential expression of virulence genes in TSB vs SSLM.
- Fig 4 compares gene expression in previous work ons skin explants and SLM. I’ve not read the details on growth of bacteria on skin explants. We do have the U of A animal farm that sells pig skin.
- Fig 5 pH and temperature on virulence gene expression.
- Fig 6 demonstrates enhanced adhesion of MRSA to human corneocytes after growth in SLM.
- Fig 7 sums up changes in TSB vs SLM, There are some changes in nucleotide synthesis gene pathways as well as fatty acid and branched chain amino acid synthesis.
Changes in the expression of potential antenna proteins
- Gene name, some of the previously mentioned regulatory genes were searched for in the supplemental data of Costa 2024.
- The Excel spreadsheet was searched for terms like”reductase” and Iron-sulfur.
- Log2 fold change is a measure used in genomics to express the change in gene expression levels between two conditions. A positive log2 fold change indicates increased expression in one condition compared to another, while a negative value indicates decreased expression. The reference is the TSB
- The smaller the p value, the higher the probability that the observed difference is not due to random change. For those who have forgotten Scientific Notation, p=6.69E-02 is 0.0669 and p=1.78E-7 is 0.00000017. The latter is more significant than the former. 2.21E-116 is an insanely small p value with many zeros to the righ of the decimal point.
Just about all of the potential Fe-S clusters and other potential antenna proteins are decreased on the transcript level. It is suggested that the non-scientist reader scroll down to the send simply taken in the size of the list.
| gene name | product | log2 Fold change | p value | p adj | TSB1 | TSB2 | TSB3 | SLM1 | SLM2 | SLM3 |
|---|---|---|---|---|---|---|---|---|---|---|
| airR | DNA-binding response regulator | 0.27 | 6.69E-02 | 0.081166 | 398 | 440 | 538 | 551 | 587 | 518 |
| airS | putative sensor histidine kinase | 0.55 | 1.78E-07 | 3.14E-07 | 647 | 698 | 685 | 956 | 1005 | 1014 |
| arcR | Crp/FNR family transcriptional regulator | -1.37 | 2.85E-27 | 9.37E-27 | 802 | 746 | 815 | 310 | 324 | 277 |
| nreB | sensor histidine kinase | -1.96 | 6.17E-79 | 4.52E-78 | 12210 | 11734 | 14353 | 3057 | 3332 | 3470 |
| nreC | DegU family transcriptional regulator | -1.78 | 4.45E-65 | 2.79E-64 | 6173 | 6653 | 7334 | 1979 | 1786 | 2111 |
| nirB | nitrite reductase [NAD(P)H], large subunit | -1.99 | 6.57E-46 | 3.12E-45 | 70968 | 60000 | 65422 | 12720 | 19624 | 17045 |
| nirD | nitrite reductase [NAD(P)H], small subunit | -2.93 | 2.21E-116 | 2.39E-115 | 12546 | 9750 | 10511 | 1209 | 1580 | 1509 |
| nirR | nitrite reductase transcriptional regulator NirR | -1.64 | 1.22E-24 | 3.72E-24 | 20965 | 16682 | 18477 | 4293 | 7219 | 6444 |
| narG | respiratory nitrate reductase, alpha subunit | -1.30 | 1.12E-20 | 3.11E-20 | 135778 | 118622 | 125422 | 39299 | 60973 | 53844 |
| narH | respiratory nitrate reductase, beta subunit | -3.38 | 1.20E-203 | 2.98E-202 | 47360 | 41132 | 45143 | 3648 | 4622 | 4595 |
| narI | respiratory nitrate reductase, gamma subunit | -2.57 | 5.07E-139 | 6.73E-138 | 7616 | 8420 | 8936 | 1316 | 1353 | 1533 |
| narJ | respiratory nitrate reductase, subunit delta | -4.61 | 9.72E-257 | 3.40E-255 | 12402 | 11491 | 12835 | 395 | 497 | 610 |
| cysJ | sulfite reductase flavoprotein | 0.58 | 1.49E-08 | 2.76E-08 | 1263 | 1440 | 1494 | 2095 | 2012 | 2156 |
| sufB | FeS assembly protein SufB | -0.66 | 6.67E-18 | 1.71E-17 | 22376 | 24342 | 23604 | 14600 | 14915 | 15007 |
| sufC | FeS assembly ATPase SufC | -0.84 | 3.00E-12 | 6.44E-12 | 8246 | 8230 | 8537 | 4416 | 4021 | 5573 |
| sufD | FeS assembly protein SufD | -1.52 | 3.76E-50 | 1.91E-49 | 20171 | 24277 | 24048 | 8571 | 7508 | 7857 |
| sufS | SufS subfamily cysteine desulfurase | -1.32 | 9.89E-51 | 5.10E-50 | 20522 | 23168 | 23063 | 9354 | 8533 | 8923 |
| sufT | DNA methyltransferase | 1.15 | 5.16E-38 | 2.13E-37 | 2924 | 3416 | 3198 | 7069 | 7065 | 7030 |
| sufU | NifU family SUF system FeS assembly protein | -1.49 | 1.69E-45 | 7.96E-45 | 8205 | 8965 | 9010 | 3075 | 2763 | 3490 |
| sdaAA | L-serine dehydratase, iron-sulfur-dependent, alpha subunit | 0.28 | 2.66E-02 | 0.033717 | 833 | 1102 | 906 | 1117 | 1171 | 1155 |
| sdaAB | L-serine dehydratase, iron-sulfur-dependent, beta subunit | 0.035 | 3.21E-02 | 0.040264 | 291 | 354 | 403 | 489 | 466 | 384 |
These changes are just a drop in the bucket of how one microorganism, Staph aureus, totally changes who it is as a bug when going from tryptic soy broth to a skin like broth under aerobic conditions! What if they had added “low O2” or a bone like broth, or some lesser known broth simulating another organ system that Staph aureus might infect? While professional scientists scramble for funds to finance mind blowing studies, perhaps citizen scientists can amuse themselves understanding how non pathogens respond to PEMF.
Parting comments..
This post makes the case that perhaps we can kill bugs as they try to revamp for different environments in our bodies. This post does not mean to suggest that PEMF can replace standard of care. This post is meant to present to a professional licensed to practice and ask, “What do you think about adding PEMF to the current treatment?
References
- Bayır E, Bilgi E, Şendemir-Ürkmez A, Hameş-Kocabaş EE. (2015) The effects of different intensities, frequencies and exposure times of extremely low-frequency electromagnetic fields on the growth of Staphylococcus aureus and Escherichia coli O157:H7. Electromagn Biol Med. 2015 Mar;34(1):14-8. Sci-Hub free paper
- Costa FG, Mills KB, Crosby HA, Horswill AR. (2024) The Staphylococcus aureus regulatory program in a human skin-like environment. mBio. 2024 May 8;15(5):e0045324. PMC free paper
- Chettouh-Hammas N, Grillon C. Physiological skin oxygen levels: An important criterion for skin cell functionality and therapeutic approaches. Free Radic Biol Med. 2024 Sep;222:259-274. free paper
- Fojt, L., Stra ˇsa´ k, L., Vetterl, V., et al. (2004). Comparison of the low-frequency magnetic field effects on bacteria Escherichia coli, Leclercia adecarboxylata and Staphylococcus aureus. Bioelectrochemistry, 63:337–341. Sci-Hub free paper
- Hall JW, Ji Y. (2013) Sensing and Adapting to Anaerobic Conditions by Staphylococcus aureus. Adv Appl Microbiol. 2013;84:1-25. Sci-Hub free paper
- Johnson RW, Sowder ME, Giaccia AJ. (2017) Hypoxia and Bone Metastatic Disease. Curr Osteoporos Rep. 2017 Aug;15(4):231-238. PMC free paper
- Loi VV, Busche T, Tedin K, Bernhardt J, Wollenhaupt J, Huyen NTT, Weise C, Kalinowski J, Wahl MC, Fulde M, Antelmann H. (2018) Redox-Sensing Under Hypochlorite Stress and Infection Conditions by the Rrf2-Family Repressor HypR in Staphylococcus aureus. Antioxid Redox Signal. 2018 Sep 1;29(7):615-636. PMC free paper
- Makhlin J, Kofman T, Borovok I, Kohler C, Engelmann S, Cohen G, Aharonowitz Y. (2007) Staphylococcus aureus ArcR controls expression of the arginine deiminase operon. J Bacteriol. 2007 Aug;189(16):5976-86. d PMC free paper
- Matl FD, Obermeier A, Zlotnyk J, Friess W, Stemberger A, Burgkart R. (2011) Augmentation of antibiotic activity by low-frequency electric and electromagnetic fields examining Staphylococcus aureus in broth media. Bioelectromagnetics. Jul;32(5):367-77. Sci-Hub free paper
- Obermeier A, Matl FD, Friess W, Stemberger A. Growth inhibition of Staphylococcus aureus induced by low-frequency electric and electromagnetic fields. Bioelectromagnetics. 2009 May;30(4):270-9. Sci-Hub free paper
- Wohlkönig A, Stalon V, Vander Wauven C. (2004) Purification of ArcR, an oxidation-sensitive regulatory protein from Bacillus licheniformis. Protein Expr Purif. 2004 Sep;37(1):32-8. PubMed
Leave a Reply to PEMF Staph aureus atopic dermatitis – PEMF BioCancel reply